Neuer Artikel: „Toward a modeling, optimization, and predictive control framework for fed-batch metabolic cybergenetics“
09.11.2023
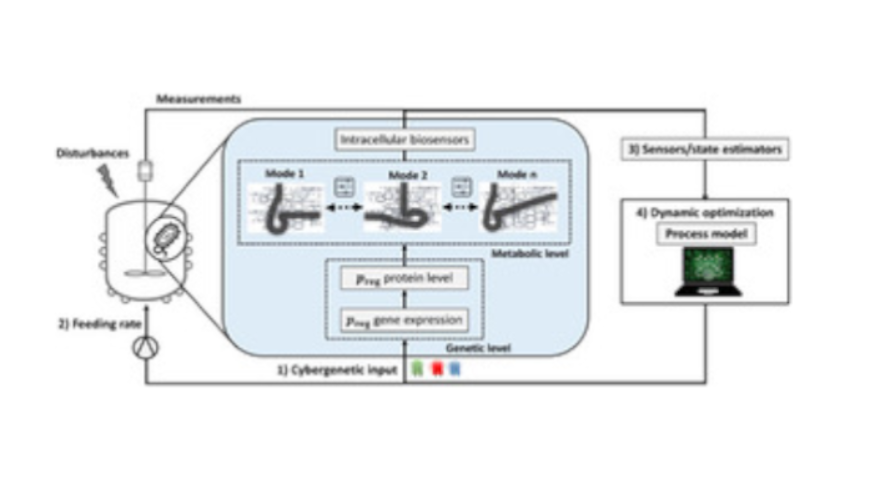
Toward a modeling, optimization, and predictive control framework for fed-batch metabolic cybergenetics
Biotechnology offers many opportunities for the sustainable manufacturing ofvaluable products. The toolbox to optimize bioprocesses includesextracellularprocess elements such as the bioreactor design and mode of operation, mediumformulation, culture conditions, feeding rates, and so on. However, these elementsare frequently insufficient for achieving optimal process performance or preciseproduct composition. One can use metabolic and genetic engineering methods foroptimization at the intracellular level. Nevertheless, those are often ofstaticnature,failing when applied todynamicprocesses or if disturbances occur. Furthermore,many bioprocesses are optimized empirically and implemented with little‐to‐nofeedback control to counteract disturbances. The concept of cybergenetics hasopened new possibilities to optimize bioprocesses by enabling online modulation ofthe gene expression of metabolism‐relevant proteins via external inputs (e.g., lightintensity in optogenetics). Here, we fuse cybergenetics with model‐based optimiza-tion and predictive control for optimizing dynamic bioprocesses. To do so, wepropose to use dynamic constraint‐based models that integrate the dynamics ofmetabolic reactions, resource allocation, and inducible gene expression. Weformulate a model‐based optimal control problem to find the optimal processinputs. Furthermore, we propose using model predictive control to addressuncertainties via online feedback. We focus on fed‐batch processes, where thesubstrate feeding rate is an additional optimization variable. As a simulation example,we show the optogenetic control of the ATPase enzyme complex for dynamicmodulation of enforced ATP wasting to adjust product yield and productivity.



